from China, for the World
for Superior Biology Services since 2000
- Products
- All Products
- Custom Services
- Catalog Products
- Innovative Systems
- Nucleic Acid Related
- Natural Compounds
- Enzymes
- POCT
- 6 POCT Platforms
- LAMP
- RPA
- CRISPR
- Freeze-Drying System
- Lateral Flow System
- DNA-Free Enzymes
- Pathogen Detection
- About
- About SBS
- Achievements
- Ecosystem
- Legal Statement
- …
- Products
- All Products
- Custom Services
- Catalog Products
- Innovative Systems
- Nucleic Acid Related
- Natural Compounds
- Enzymes
- POCT
- 6 POCT Platforms
- LAMP
- RPA
- CRISPR
- Freeze-Drying System
- Lateral Flow System
- DNA-Free Enzymes
- Pathogen Detection
- About
- About SBS
- Achievements
- Ecosystem
- Legal Statement
from China, for the World
for Superior Biology Services since 2000
- Products
- All Products
- Custom Services
- Catalog Products
- Innovative Systems
- Nucleic Acid Related
- Natural Compounds
- Enzymes
- POCT
- 6 POCT Platforms
- LAMP
- RPA
- CRISPR
- Freeze-Drying System
- Lateral Flow System
- DNA-Free Enzymes
- Pathogen Detection
- About
- About SBS
- Achievements
- Ecosystem
- Legal Statement
- …
- Products
- All Products
- Custom Services
- Catalog Products
- Innovative Systems
- Nucleic Acid Related
- Natural Compounds
- Enzymes
- POCT
- 6 POCT Platforms
- LAMP
- RPA
- CRISPR
- Freeze-Drying System
- Lateral Flow System
- DNA-Free Enzymes
- Pathogen Detection
- About
- About SBS
- Achievements
- Ecosystem
- Legal Statement
CRISPR Gene Editing
At SBS Genetech, we are at the forefront of offering tools for CRISPR Gene Editing, from basic materials to complete solutions
What is CRISPR?
A brief introduction of CRISPR gene editing system

Clustered Regularly Interspaced Short Palindromic Repeat (CRISPR) is a bacterial defense system that forms the basis for CRISPR/Cas gene-editing technology. This new gene-editing technology is now becoming a more efficient and customizable alternative to other existing tools.
CRISPR system contains two components: a CRISPR-associated endonuclease (Cas nuclease) and a single guide RNA (sgRNA). Cas protein snips through DNA like a pair of molecular scissors, and sgRNA directs Cas protein to a specific site of DNA to make the cut. The protospacer adjacent motif (PAM) is a short DNA sequence following the target cutting site. The PAM is also a prerequisite for a Cas nuclease to cut.
Cas Nucleases
Choose the most suitable molecular scissor for your research
Cas13 recognizes and cleaves target RNA under the guidance of guide RNA, its collateral cleavage activity is activated, which can efficiently cleave non-specific single-stranded RNA (ssRNA). The lyophilized version of Cas13a can be transported at room temperature, saving the high cost of dry ice transportation.
Cas12a (Cpf1) belongs to class II and type VI CRISPR system effector proteins, and is an endonuclease that binds to and cleavages specific sites of target DNA under the guidance of single-stranded guide RNA. The lyophilized version of Cas12a can be transported at room temperature, saving the high cost of dry ice transportation.
Cas9 nuclease can form ribonucleoprotein (RNP) complex with the single guide RNA (sgRNA) component of the CRISPR/Cas9 system, inducing site-specific DNA double-stranded breaks. The PAM sequence is NGG.
dCas9 protein is a ribozyme deficient Cas9 protein, which does not have sgRNA guided DNA strand cleavage activity, but retains target DNA binding activity, regulating the gene expression.
AapCas12b is an RNA-mediated endonuclease that binds to and cleaves specific sites of target DNA under the guidance of single-stranded guide RNA.
spCas9-NG is an RNA-mediated nuclease, which can catalyze the specific site cleavage of double-stranded DNA. It can recognize the NG PAM sequence.
When LwCas13a recognizes and cleaves the target RNA under the guidance of guide RNA, its "accessory cleavage" activity is activated, which can efficiently cleave non-specific single-stranded RNA (ssRNA) in the reaction system.
SpRYCas9 is an RNA-mediated nuclease, which can catalyze the specific site cleavage of double-stranded DNA. It can recognize the NNN (NRN > NYN) PAM sequence, which almost gets rid of the restriction of the PAM.
LbaCas12a (Cpf1) has a RuvC endonuclease domain similar to Cas9 but does not have the HNH endonuclease domain. LbaCas12a is not only smaller than Cas9, but also requires a smaller RNA (nearly half of Cas9's total sgRNA), which is very helpful for genome editing.
Compared with other Cas proteins, the molecular weight of Cas14 protein is generally smaller (400-700aa). Similar to Cas12, Cas14a1 can also bind the target nucleic acid and activate its ssDNA trans cleavage activity.
sgRNAs
A programmable GPS from different sources
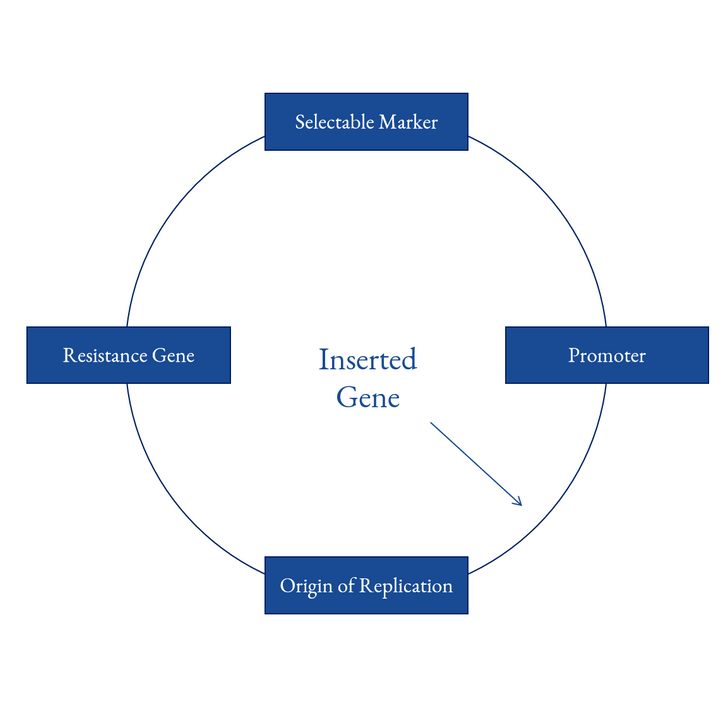
 1st Generation
1st GenerationThe sgRNA sequence is first cloned into a plasmid vector and then introduced into cells by transfection, which is suitable for High-Throughput Gene Editing. This is the most original method, which requires more than a week for the preparation before the gene-editing experiment.

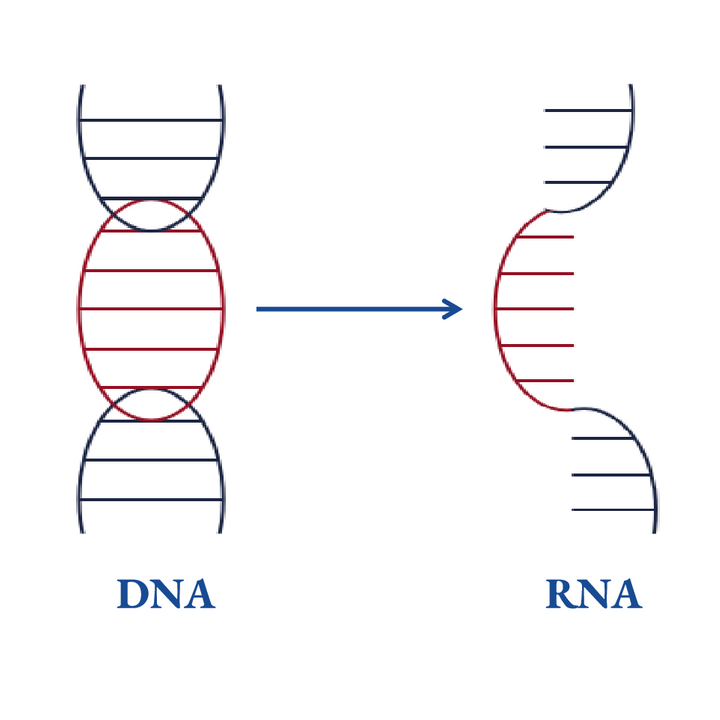
2nd Generation
The sgRNA is first transcribed from a DNA template by RNA polymerase. Additional purification is then required before the experiment. Generally, making an In Vitro-transcribed (IVT) sgRNA takes around 3 days.

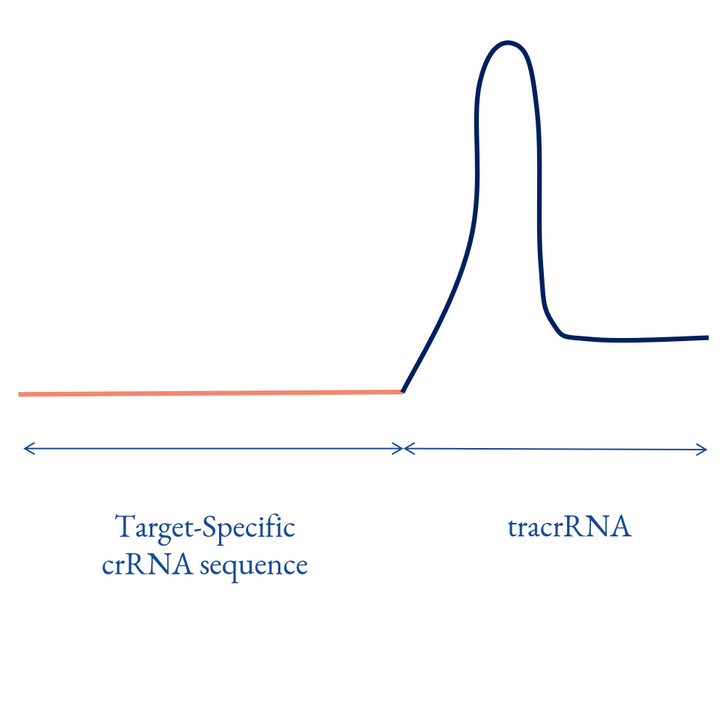
3rd Generation
The sgRNA is directly synthesized via a chemical approach. Research shows that synthetic sgRNA has more consistent editing efficiencies and lower off-target effects compared with plasmid and IVT sgRNAs.
Ready for this new experience?
Custom Services
 Synthetic pegRNA (Prime Editing guide RNA)
Synthetic pegRNA (Prime Editing guide RNA) Synthetic Cas12a (cpf1) crRNAs
Synthetic Cas12a (cpf1) crRNAs Synthetic Ultrapure sgRNAs (HPLC)
Synthetic Ultrapure sgRNAs (HPLC) Synthetic sgRNAs
Synthetic sgRNAs Human Epigenetic CRISPR Knockout Library
Human Epigenetic CRISPR Knockout Library Human UBDUB CRISPR Knockout Library
Human UBDUB CRISPR Knockout Library Pig CRISPR Knockout Pooled Library
Pig CRISPR Knockout Pooled Library Green Monkey CRISPR Knockout Pooled Library
Green Monkey CRISPR Knockout Pooled Library Mouse CRISPR Knockout Pooled Library (Single Plasmid System A)
Mouse CRISPR Knockout Pooled Library (Single Plasmid System A) Mouse CRISPR Knockout Pooled Library (Single Plasmid System B)
Mouse CRISPR Knockout Pooled Library (Single Plasmid System B) Mouse CRISPR Knockout Pooled Library (Single Plasmid System A+B)
Mouse CRISPR Knockout Pooled Library (Single Plasmid System A+B) Mouse CRISPR Activation Library (SAM - 3 Plasmid System)
Mouse CRISPR Activation Library (SAM - 3 Plasmid System) Mouse CRISPR Knockout Pooled Library (Dual Plasmid System A)
Mouse CRISPR Knockout Pooled Library (Dual Plasmid System A) Human CRISPR Knockout Pooled Library (Single Plasmid System A)
Human CRISPR Knockout Pooled Library (Single Plasmid System A) Human CRISPR Knockout Pooled Library (Single Plasmid System B)
Human CRISPR Knockout Pooled Library (Single Plasmid System B) Human CRISPR Whole Genome Knockout Library (Single Vector System A+B)
Human CRISPR Whole Genome Knockout Library (Single Vector System A+B) Human CRISPR Whole Genome Knockout Library (Dual Vector System A)
Human CRISPR Whole Genome Knockout Library (Dual Vector System A) Human CRISPR Whole Genome Knockout Library (Dual Vector System B)
Human CRISPR Whole Genome Knockout Library (Dual Vector System B) Human CRISPR Whole Genome Knockout Library (Brunello)
Human CRISPR Whole Genome Knockout Library (Brunello) Human CRISPR Whole Genome Activation Library
Human CRISPR Whole Genome Activation LibraryProducts
 Buy nowdCas9 NLS (Lyophilized)$240.00 - $1,280.00$1,600.00
Buy nowdCas9 NLS (Lyophilized)$240.00 - $1,280.00$1,600.00 Buy nowEiCsm6 (Lyophilized)$1,386.00 - $3,850.00$5,500.00
Buy nowEiCsm6 (Lyophilized)$1,386.00 - $3,850.00$5,500.00 Buy nowLwaCas13a Nuclease (Lyophilized)$350.00 - $1,610.00
Buy nowLwaCas13a Nuclease (Lyophilized)$350.00 - $1,610.00 Buy nowBrCas12a Nuclease (Lyophilized)$771.00 - $4,950.00$5,500.00
Buy nowBrCas12a Nuclease (Lyophilized)$771.00 - $4,950.00$5,500.00 Buy nowAsCas12a Nuclease (Lyophilized)$308.00 - $1,463.00$2,090.00
Buy nowAsCas12a Nuclease (Lyophilized)$308.00 - $1,463.00$2,090.00More Information
August 2, 2025Read more...🌟 A Benchmark in Integrated Biotechnology In today’s fast-evolving landscape of molecular...January 20, 2024Read more...In this era of rapid technological advancement, CRISPR diagnosis has emerged as a captivating...January 10, 2024Read more...Since the introduction of the groundbreaking CRISPR/Cas technology, it has received widespread...Read more...The primary objective of effectively managing infectious diseases and epidemics is the prompt and...Published Papers
Representative Publications Using SBS Genetech CRISPR Gene Editing Products
Kershanskaya, O.I., Yessenbaeva, G.L., Nelidova, D.S., Karabekova, A.N. & Sadullaeva, Z.N. (2022) CRISPR/Cas genome editing perspectives for barley breeding. Physiologia Plantarum, 174( 3), e13686.
Wang K, Huang W, Chen R, Lin P, Zhang T, Ni YF, Li H, Wu J, Sun XX, Geng JJ, Zhu YM, Nan G, Zhang W, Chen X, Zhu P, Bian H, Chen ZN. (2021) Di-methylation of CD147-K234 Promotes the Progression of NSCLC by Enhancing Lactate Export. Cell Metabolism.
Ibañez Oliver, A. M. (2021). Desarrollo de un "pipeline" de trabajo para secuenciación dirigida por CRISPR-Cas9 (Trabajo final de carrera). Universidad ORT Uruguay, Facultad de Ingeniería.
Trofimenko E, Grasso G, Heulot M, Chevalier N, Deriu MA, Dubuis G, Arribat Y, Serulla M, Michel S, Vantomme G, Ory F, Dam LC, Puyal J, Amati F, Lüthi A, Danani A, Widmann C. (2021) Genetic, cellular, and structural characterization of the membrane potential-dependent cell-penetrating peptide translocation pore. Elife.
Zhang, R., Du, J., Zhao, X., Wei, L. and Zhao, Z. (2021), Regulation of circadian behavioural output via clock-responsive miR-276b. Insect Mol Biol, 30: 81-89.
Wang, G.-F., Niu, X., Liu, H., Dong, Q., Yao, Y., Wang, D., Liu, X. and Cao, C. (2021), c-Abl kinase regulates cell proliferation and ionizing radiation-induced G2/M arrest via phosphorylation of FHL2. FEBS Open Bio, 11: 1731-1738.
Zhong X, Zhang W, Sun T. (2019) DDR1 promotes breast tumor growth by suppressing antitumor immunity. Oncol Rep.
He Y, Wang M, Liu M, Huang L, Liu C, Zhang X, Yi H, Cheng A, Zhu D, Yang Q, Wu Y, Zhao X, Chen S, Jia R, Zhang S, Liu Y, Yu Y, Zhang L. (2018) Cas1 and Cas2 From the Type II-C CRISPR-Cas System of Riemerella anatipestifer Are Required for Spacer Acquisition. Front Cell Infect Microbiol.
Still Can’t Find What You Need?
SBS Genetech EcosystemTM helps you to find life science products from more Chinese suppliers

- ContactFor more information, please fill out the provided form or contact us directly by E-mail
SBS Genetech © Copyright 2000-2026
from China, for the World
for Superior Biology Services since 2000